-
Mitochondrial Complex I Deficiency, Nuclear Type 23
Omim
Clinical Features Ostergaard et al. (2011) reported a 10-year-old girl, born of consanguineous Pakistani parents, with mitochondrial complex I deficiency manifesting as Leigh syndrome (see 256000). The patient had delayed motor development, with walking at age 20 months. ... Molecular Genetics In a 10-year-old girl, born of consanguineous Pakistani parents, with complex I deficiency manifesting as Leigh syndrome, Ostergaard et al. (2011) identified a homozygous nonsense mutation in the NDUFA12 gene (614530.0001). ... INHERITANCE - Autosomal recessive GROWTH Other - Poor overall growth SKELETAL Spine - Scoliosis SKIN, NAILS, & HAIR Hair - Hypertrichosis MUSCLE, SOFT TISSUES - Hypotonia - Muscle atrophy NEUROLOGIC Central Nervous System - Delayed motor development - Delayed walking - Progressive loss of motor abilities - Dystonia - Learning difficulties - White matter abnormalities consistent with Leigh syndrome LABORATORY ABNORMALITIES - Mitochondrial complex I deficiency in various tissues MISCELLANEOUS - Onset in early childhood - Progressive disorder - One patient, born of consanguineous Pakistani parents, has been reported (last curated January 2019) MOLECULAR BASIS - Caused by mutation in the NADH-ubiquinone oxidoreductase subunit A12 gene (NDUFA12, 614530.0001 ▲ Close
-
Athymhormia
Wikipedia
See also [ edit ] Aboulia Amotivational syndrome Anhedonia Athymhormic syndrome Avolition References [ edit ] Further reading [ edit ] Patrick Verstichel and Pascale Larrouy. ... "[Loss of vitality, of interest and of the affect (athymhormia syndrome) in lacunar lesions of the corpus striatum]".
-
Glioblastoma
Gard
In most cases, the exact underlying cause is unknown; however, they can rarely occur in people with certain genetic syndromes such as neurofibromatosis type 1 , Turcot syndrome and Li Fraumeni syndrome .EGFR, PTEN, MTOR, H3-3A, FN1, NF1, HIF1A, MDM4, CXCL8, CTSB, MET, TP53, PROM1, MGMT, NOTCH1, TNFSF10, IL1B, PDGFRA, MMP2, MMP9, BMI1, VEGFA, CHI3L1, MYC, EGF, IL2, NOTCH2, H2AX, GDNF, PTK2, PRKN, TP53BP1, CDK6, WT1, CTSK, TGM2, NDRG1, APC, RECK, JAG1, H3C2, HEY1, LZTR1, RUNX1, BRD2, TES, CD9, DAXX, PML, RUNX3, BHLHE40, PIM1, FAT1, GSTT1, SRRT, BRD4, POLK, SUZ12, SEPTIN14, NOTCH3, FOSL2, HES1, DUSP6, BCHE, LTBP4, JAG2, IFNA2, CSTA, GRIK2, ZBTB7A, TRMT11, CSTB, TMEM135, MACIR, CTNND2, DEUP1, SLC22A10, LRRC59, PTGS2, NCOR1, CCNH, BAG3, CPEB1, ADGRL4, SERPING1, PTGS1, PIK3CA, MYCN, IL16, ATM, FGFR1, BRAF, IDH1, MSH2, RTEL1, PMS2, PIK3R1, NRAS, CCDC26, FBXW7, POLE, HRAS, DNMT3A, SSBP2, POLR3B, CDKN2B-AS1, CDKN1B, RAVER2, IFNB1, SLC12A9, IFNG, TNF, CDKN1A, MIB1, HSPA4, IGF1, HEATR3, CD34, CDKN2A, CDK4, IDH2, PDCD1, HSPB1, FGF2, IGFBP2, CASP8, SERPINE1, TNC, HSP90AA1, HTC2, CASR, CAV1, GFAP, CDKN2B, AHSA1, PRKAR1A, PRKCA, PRMT5, YAP1, MYCNOS, OLIG2, TRIM13, ROS1, AIMP2, HDAC6, NR1I3, MAPK1, MAPK3, MAPK8, GDF15, ABCG2, GRAP2, ERBB2, HMGA2, EPHB2, HGF, PTN, E2F1, MBD2, H3-3B, TNFRSF10B, EPHA2, RAC1, MSH6, CIB1, CXCR4, CD274, CXADR, CXCL12, SETD2, PDGFRB, PTK2B, PDCD4, CCL2, POLDIP2, RNF19A, VIM, SLC16A8, ABCB1, RHBDF1, PIK3CB, F3, PIK3CD, PIK3CG, TBC1D9, PKM, PLAU, CREBBP, CRK, MAPK14, CSF2, POU5F1, PPARA, EZH2, NES, CTNNB1, WNT5A, COL11A2, CD44, CASP3, FOXM1, AKT2, IL13, AZIN2, AKT1, MIR221, ATRX, COX2, IL6, MIR21, KDR, STAT1, TGFB1, BAX, TGFB2, SPG7, IL13RA2, MIR222, TERT, GADL1, GJA1, ZEB1, GABPA, MCL1, ALK, SPP1, MIR34A, C11orf65, MDM2, GLI1, BCL2, CCND1, NFE2L2, MIR10B, KIT, MMP14, TSPO, ACTB, MIR137, ABCC1, LGALS1, MKI67, RTEL1-TNFRSF6B, COL18A1, MIR4300HG, ITGA5, ARR3, LOC110806263, H3P10, MIR145, HOTAIR, ANGPT1, AQP4, POU5F1P4, STAT3, CA9, SOX2, MSH3, PARP1, CXADRP1, LIG4, MIR203A, MSI1, POU5F1P3, ANGPT2, APOE, AURKA, MALAT1, ANXA2, APEX1, MLH1, S100A4, CCN2, STAT5B, PPARG, AXL, FLT1, IL4, RELA, RAD51, WNK1, MTCO2P12, NANOG, CCK, IL1A, THBS1, ACKR3, NRP1, HSPB3, AKT3, KLF4, DNMT1, ITGAV, HDAC9, CDK1, DMBT1, TNFRSF12A, SLC2A1, HSPB2, CD47, FOXO3, TMED7-TICAM2, HMGB1, KLRC4-KLRK1, SYT1, ATF5, BECN1, PTPRZ1, XRCC6P5, KLRK1, LILRB1, PLK1, GORASP1, FAS, WEE1, PTPN11, MAP2K7, MIR210, AHR, EPO, RALBP1, TACC3, GSK3B, SIRT1, NFKB1, FOLH1, TMED7, IDO1, TICAM2, PCNA, GLS2, PAX6, OSM, IFNA1, IFNA13, MXI1, IGF1R, CCNB1, TLR4, EPAS1, CCN1, TP73, DECR1, ELAVL2, CDK2, MIR7-1, FOXO1, HSPA5, H3P23, CASP9, ZNRD2, POSTN, TIMP2, PRNP, EDNRA, PSMD9, BRCA1, MIR340, TIMP1, TGFA, SMAD2, TXN, ADM, MIR146B, IFI27, SOCS3, EGR1, NR1I2, SLC16A1, SOX9, HMGA1, CRYAB, DCTN6, MIF, XIAP, KDM1A, CTSL, CSPG4, VDAC1, SMN1, CEBPB, CTLA4, SMN2, TAZ, CXCR6, SMUG1, MIR139, SESN2, PECAM1, ARHGEF7, SPHK1, SQSTM1, KRAS, IL10, ADARB1, CMA1, FASLG, HDAC1, MIR181D, POLD1, SLC7A5, SOD1, ATN1, GHRH, PLG, BCL2L12, CREB1, GSN, KLF6, GSTP1, NTRK1, GOLPH3, MIR182, SOAT1, TACR1, FGFR3, EDN1, PLAUR, NOS2, EPHA1, MRC1, CD276, RIPK1, LGALS3, EPHA3, GLS, STAT5A, HMOX1, MIR7-3, SNAI2, PRKDC, MIR148A, ROCK1, TFPI2, NUDT21, ILK, BCL2L11, LINC01194, ALDH1A3, MIR7-2, HSPA1B, MCTS1, JUN, CCNO, ESR2, CDC42, MCAT, HSPA1A, CLIC1, PRKAA2, APP, TWIST1, MIR338, SMAD3, TTK, PAEP, MIR326, MIR29A, FABP7, RRM2, SRC, TAC1, SSTR4, SREBF1, SOD2, GPR42, UQCRFS1, REST, TIMP3, F2R, GLI2, RPS27, VEGFC, FOSB, CCL5, TFRC, SLC12A2, SPARC, ERCC1, ERBB3, TOP1, TOP2A, SNAI1, SAI1, FOS, WWTR1, IDUA, ICAM1, IGFBP3, MIR195, MFAP1, NDRG2, NTSR1, NUP98, APRT, TRPM7, CDC20, LIN28A, GDE1, MIR296, HSF1, MDK, NOX4, ADGRE2, MIR31, EEF2K, NT5E, CALR, HPGDS, MIR155, ASL, ASAH1, MIR143, STS, MPEG1, PDIK1L, GSC, MIR146A, PRRT2, MMP1, DNER, MTDH, NF2, BRS3, BSG, WNT3A, BUB1B, IL4R, NXT1, PDGFB, ALOX5, MRPL28, EBI3, PRKCI, L1CAM, COMMD3-BMI1, LRIG2, ATG5, H3P9, HLA-A, HK2, CD163, LAMC2, LPAR2, SLC16A3, SLC16A4, IL18R1, ABCC3, PVR, ELK1, RAF1, BBC3, TUBB3, PRKAB1, PDPN, LAT, LRIG1, CCR5, PFKFB3, PGR, ALB, COX8A, MAOB, FSTL1, MIR134, GLIPR1, SUB1, STAG2, ADRA2B, ADRA1A, PRKAA1, LEP, JUND, JAK2, GLB1, LIMK1, MST1, JUNB, RBL2, MTAP, NBN, PTBP1, GRN, GRIA1, HMMR, PRKCB, S100B, PTPA, HOXA9, SATB1, PDK1, SGK1, SH3GL2, SKP2, GLUL, PRDX1, P4HB, RB1, IGF2, NTS, NFKBIA, SLC22A3, NEFL, IL17A, SMARCB1, NCL, CXCL10, NCAM1, LGI1, LDLR, SOS1, TLR9, FOXP3, LEF1, PHF20, TRPV2, CD151, ADGRE5, CD70, CD68, TIGAR, CD40, RMDN3, TNFRSF19, NLRP2, PBK, AJAP1, METTL3, CEACAM5, CHEK1, SLC7A11, KCNH4, MELK, USP15, NAMPT, MERTK, TRIM3, CX3CR1, SPINT2, HPSE, DSTN, CSF3, WIF1, CHEK2, MMRN1, NT5C2, PHLDB1, SPHK2, SEMA6A, IKBKE, MIR223, ASCL1, MIR144, ARRB1, MIR149, MIR183, AR, AQP1, MIR29B1, GAS5, MIR485, MIR504, ADARB2, MNX1-AS1, MAGED4, LINC01672, SERPINA3, MIR141, MIR100, ADGRB1, CAVIN1, LRRC4, TRPM8, CAPG, SLC52A2, TET1, CALCR, DDR1, ARHGAP24, BUB1, KLF9, KCNH8, CD109, BDNF, PWAR1, BCAT1, HDAC4, NOP53, TUSC3, FCGR2B, TEP1, KAT2B, FOXG1, TNFAIP3, CFLAR, TNFRSF1A, LGR5, EFEMP1, FLT3LG, TYMS, EMP2, EMP3, VTN, XIST, USP9X, TFEB, DUSP1, ESR1, ADAM17, SP1, DRD2, STAT6, MIR29B2, USP1, AGAP2, PRAP1, LRIG3, TRPV1, VCAM1, NEDD9, STC1, MIR454, CBLL2, LIF, HOXA11-AS, ASPM, ATP23, ORAI1, NFATC2, MAGED4B, SEMA3B, MUL1, WLS, ZFP36, S100A1, XRCC3, TRIM8, LOX, S100A9, TCHP, BEX2, S100A10, ST13, HAVCR2, TEAD1, LRP1, MIR17HG, SLC9A9, MIR18A, MIR181C, MARCKS, MIR451A, MIR186, TGFBI, MIR370, MIR330, MIR197, MIR20A, MIR205, MIR219A1, MIR135B, MIR224, MIR299, MIR154, MIR152, MMP3, MIR125A, MACC1, MTHFR, SKI, MIR106A, MIR10A, MSN, MIR126, TEK, FTO, MIR142, TRPM2, SLC2A3, TRAF2, MMP12, TSC1, GGPS1, NPM1, MPRIP, PODXL, H3C7, PDCD10, OIP5, APLN, KDM6B, CIC, PIN1, DICER1, RPS6, SIRT3, MBTPS1, PHF3, CBX7, CDK20, PGF, CORO1C, TRADD, FAM107A, OSMR, BUB3, IL24, MAPK9, WTAP, PROX1, KMT2B, LGMN, KLK6, ZEB2, AIM2, MVP, ABCB6, NTN1, COX5A, SEMA3C, SLC9A3R1, AIMP1, KHDRBS1, AURKB, DKK3, PTPRD, SGSM3, XAF1, SLC22A18, ALKBH5, H3C4, H3C1, FZD7, RNASE3, TAM, LINC00470, YEATS4, ROBO1, GOPC, KMT2D, MRTFA, CXCL16, NTN4, BCAR1, MARCKSL1, H3C3, DLL4, PYCARD, ERRFI1, EIF3B, SOX4, IL22, SOCS1, ASCC1, EXOSC3, ANGPTL4, PDGFA, H3C10, H3C12, RNF138, MPC1, H3C8, WWOX, PAK1, H3C11, H3C6, A1BG, H3P40, VPS51, BCL2L2, BIRC3, FASN, IL2RA, BIRC5, BCL6, FPR1, ITGA7, AIF1, BNIP3, FUS, APAF1, IKBKB, DOCK1, CSF1R, IGFBP7, DRD5, DSC3, SEPTIN7, GBP1, HBEGF, CTSD, ANXA7, GLI3, HLA-G, INPPL1, CCND3, FGFR2, ING1, CCND2, DGKA, DCN, VEGFD, CYP27B1, CYP19A1, ANXA5, ITGA6, FKBP4, BMP4, BMP7, GADD45A, ANG, DDIT3, FDPS, IRS1, ETS1, GAPDH, CCNE1, ERBB4, CALM2, KPNA2, EMX2, CALM3, CNR1, EMP1, MARK2, ID4, ENG, EIF4EBP1, BTK, CDH11, CPOX, CALM1, ATR, ADAM10, COL3A1, GRIA2, ASS1, ESRRB, RND3, AXIN2, LRG1, AXIN1, SNORD35B, THEM4, SPZ1, FZD6, OLIG1, CRISP2, CCT3, TRAF6, SPPL3, RNF135, MINDY4, TPM3, MAGT1, TMPRSS13, ELF4, NMS, BMP2, IMMP2L, FGF13, RAD54L, FSD1L, EPHA4, SNORD14B, MIA, XPO1, WIPF1, BRCA2, CLDN5, PYGO2, BTC, LMLN, ST8SIA4, F10, XRCC1, ETS2, DAP3, YY1, SNORD14E, SNORD14D, VWF, EZR, EPS8, UMOD, TYR, SNORD14C, UCN, USP5, UGCG, FOSL1, ADAM12, FAAH, MCM3AP-AS1, FAP, VCP, RIOX2, VDR, ERN1, SUMO1, PPP1R13L, DCBLD2, TM7SF2, SFRP4, GNRHR, MGR3, SHC1, SIAH1, GMFB, SLC3A2, SLC5A5, SBF2-AS1, SLC9A1, SHPRH, GLUD1, PTF1A, SMARCA1, CTAG1A, SNCA, FSCN1, FOXK1, GFPT1, SOX3, GATA6, SELP, FREM2, ACTBL2, S100A11, ATP1B2, RPS6KA3, RPS6KB1, RPS15A, MIRLET7D, RRAD, MIRLET7B, RSU1, CXCR3, GPI, CXCL11, SAA1, ATP7B, TSPAN31, SAT1, GSTK1, CCL3, CCL4, CCL8, CCL11, GATA4, SOX11, GATA3, TFF3, TDO2, PRDX2, FMOD, FLT4, TERF1, PRIMA1, TF, TFAM, NR2F2, FKBP5, TDGF1, GPR65, TGFBR2, FHL2, FHIT, TIMP4, TLR1, TLR3, NR2E1, SLCO6A1, TDGF1P3, TRBV20OR9-2, PRSS55, BCL2L1, SPAST, GAS1, SPINT1, TET3, SRPK1, GAB1, XRCC6, SLC37A4, G6PC, FUT4, TCF7, STIM1, BCL3, SULT1A1, SYP, FRZB, TAPBP, RMDN2, CNTN2, HNF1A, TGFBR1, TNFRSF10A, PIK3R3, SYBU, METAP2, SLC27A3, RIPK3, TENT4A, ADAMTS5, PTPRT, NLGN3, PTP4A3, CD63, NANS, SHC3, ZHX1, KLF8, VSIG4, ACOT7, INPP5K, TUSC2, CD74, CRP, RAB3GAP1, SCAP, TET2, STIP1, CTAG1B, CD36, POMGNT1, PDLIM5, MEG3, IGF2BP2, CD86, IMP3, NEIL3, EBP, NFAT5, PLK4, ENTPD5, CEP55, PLK2, NEU3, PID1, FGL2, MKS1, ELOVL2, MALT1, ENTPD1, SNW1, DKK1, PTGR1, SH3BP4, CDR1, ERVW-1, PATZ1, CLEC5A, BACE1, CCR3, NUP62, RMC1, UHRF1, CEBPD, SMARCAL1, NOB1, CLN3, POLL, ARHGEF26, CHTOP, CLU, CHD4, NUPR1, FOXP1, CNP, CNTF, SIRT6, CDH13, DCDC2, MYT1L, CDH1, AMOTL2, CDH2, CRLF3, CPE, PLEKHM2, PHLPP1, NUSAP1, MACF1, DCTN4, ING4, CDK5, RMDN1, GAL, CDK7, CDK9, CDKN3, DELEC1, CHD7, AGR2, EIF5A, HIF1AN, TYMP, CAMKMT, E2F5, E2F3, DYRK1A, CLDN1, FSD1, INA, DUSP2, NEURL1, DTX1, ARHGEF2, LRRFIP1, NOLC1, CAPN2, MSC, CAST, NDRG4, ADGRG1, ASIC3, DPYSL2, L2HGDH, ECT2, HDAC3, PDCD1LG2, PSMG1, NUMB, EIF4E, PEA15, EFNB3, CA12, ACTL8, TNFSF13, ADAM9, TNFRSF6B, S1PR1, TNFRSF11A, TNFRSF10C, PPP1R2C, MIR122, CPEB4, TRIM24, EDNRB, SPHKAP, IQGAP1, SLIT2, DNM2, HHIP, DLEU1, TRAP1, OPTN, G3BP1, DBI, CEBPZ, TENM1, PDXP, DAPK1, PAK4, SLC2A4RG, SALL4, TUBA1B, IRF9, DIABLO, CD8A, VAV3, CYP17A1, VAT1, VPS11, ATG7, DPP3, DCC, DLD, BCAN, FHL5, CASP7, ROCK2, NAPSA, CAT, TBPL1, CXCL14, NQO1, DHCR24, RUNX2, SCO2, DES, PIEZO1, NGB, CIP2A, ARHGAP21, DCX, PREX1, DCT, KRIT1, RPL36A, TRIP6, CKB, LOXL1-AS1, HNRNPA2B1, ALDH3A1, MIR96, ITGAM, MIR99A, MIR1275, ALDH1A1, PPP1R1A, PIM3, MMP15, MMP16, PPIA, HOXA@, CD200, MPL, MIR34C, ALCAM, HOXA10, HOXA11, ITGA2, AHSG, MIR328, TNRC6C-AS1, MSMB, MIR331, HOXB2, AGAP2-AS1, PLEK, HOXB3, APLNR, ALPI, ALPP, PTPN12, ITIH4, MBP, MC4R, PTPN2, KCNH2, CD46, MIR200C, MIR204, BCL2L2-PABPN1, HK1, PTCH1, ASIC1, ME1, HLA-C, MEFV, HOTAIRM1, MIR3189, HLA-DPB1, PROS1, MIR23A, MIR27A, MGP, HLA-DRB1, MAP2K6, MAP2K1, AMFR, MIR29C, ITGB4, MIR30A, MIR30B, PRKD1, MIR339, AGT, PKD1, NFIA, NEDD8, MIR663A, NELL1, PDE1C, ADCYAP1R1, NEO1, NEU1, NFATC1, MIR503, PAX5, CXCR2, HTR7, IAPP, MIR486-1, MRPL58, MIR940, POTEM, ID1, ID2, CXCR1, MIR622, NGF, NNMT, NOS1, UCA1, H3P38, IGFBP4, IGFBP5, YBX1, RBPJ, ACTN4, PRMT1, NAP1L1, MYT1, MIR196B, HOXB7, HOXB8, MIR376A1, HOXC6, MIR378A, HOXC10, IRF3, MIR873, NUDT1, MIR425, MTTP, MTR, SERPINA1, PHB, MIR744, PGK1, HP, ACTC1, ACTG1, PFKM, MIR20B, ACTG2, MIR431, AEBP1, HPGD, ENPP1, MIR409, HPR, HELLS, POTEKP, RASGRF1, MIR132, GTF2H1, RAC3, SMAD4, RFX1, LBR, TRIM27, LCN2, GSR, MIR17, RFC4, ARG2, RET, RAP1B, RHOA, LDHA, AREG, LYN, ABCA1, GRP, RECQL, ABL1, LEPR, PLAAT4, PDIA3, KPNB1, MNS16A, ARSA, LAG3, GRPR, KDM5A, SNORD15A, ACACA, MIR191, AQP9, MIR127, NECTIN1, MIR190A, ARAF, ROBO3, CHRM3, SNHG9, CDH19, LUZP6, AVEN, TRBV9, MIR758, SLC17A7, FNDC3B, NKIRAS2, MIR671, AKR1B10, CCNG2, TRIB2, CCT6A, CCR2, CEACAM7, NBAT1, MIR579, SCYL1, SNORD118, AICDA, VANGL2, TP73-AS1, MCOLN1, ZMIZ1, CREB3L2, CEACAM3, D2HGDH, SLURP1, CHAT, PLXDC1, CD248, POTEF, RAB25, CHPT1, PMEPA1, BIRC6, CDX2, EIF5A2, AAVS1, BCCIP, MIR877, MIR577, NIBAN2, NT5DC2, PRMT8, ICOS, CDSN, PARD3, MIR574, GKN1, LRRC8A, ABO, PDGFC, MRPL41, CD3D, POLE4, DHX33, MUC13, PNO1, BDH2, HIPK2, DPYSL5, RAD18, CFTR, MIMT1, GINS3, CD1C, CEBPA, AGO2, MIR300, MIR876, CTNNA3, CHUK, RETN, SAR1A, RABGEF1, MIR582, GPR158, PJA1, MIR613, RNF123, SERP1, MIR627, H3P5, MIIP, RINT1, NIF3L1, CHRM2, CBL, SAV1, SOX17, TGIF2, NFKBIZ, HPLH1, RND1, ADAMDEC1, CASP5, ARHGEF28, ELMO2, MIR603, KIF9, MIR590, IL17B, IFIH1, TINAGL1, MIR595, MIR598, TNMD, TNN, MIR605, TENT4B, SAMSN1, CAV2, MIR610, NPAS3, ERVK-6, LGR6, RXFP1, SERPINH1, MIR652, ANKRD36B, MIR648, CD8B, TSHZ3, CBS, ANGPTL3, MIR92B, PCDH10, MARK4, RPS6KA6, MIR656, CCNA2, CASP1, SRGAP1, UBE2S, TNS3, MIR589, MIR641, RBMS3, CBR1, IL21, MMP25, VPS4A, H3P42, IL22RA1, DLGAP3, PLEKHB1, SIGLEC9, MIR630, H3P30, BHLHE22, PTBP2, CASP2, C6orf47, POLD4, SUGP1, STIM2, CSAD, RN7SL263P, PCAT1, CD58, CCHCR1, KRT20, ATAD2B, EGFL7, CDK3, MIR3666, HERC5, RIN2, MIR3908, TERF2IP, ERVK-18, GAPLINC, TREM2, ACP1, HIF1A-AS1, MZB1, LOC100505909, CD69, JPT1, EGLN1, MIR4284, PPP1R12C, DYM, NDUFA13, AHI1, MIR1238, TRIT1, MIR1179, MIR1908, LINC-ROR, ACP3, ERVK-25, ADIPOR1, HOXB-AS1, ERVK-24, TRIM44, IL17RD, CD40LG, OTUD4, DDX56, GSAP, DUOX1, MIR1246, S1PR5, STYXL1, RASD1, TRMT6, ASIC2, MIR4443, CHST15, ACAT1, CDC25A, MIR4758, CDC25B, HDAC7, ANGPT4, NCKIPSD, MIR4484, LCMT1, CDC25C, ERVK-10, ERVK-9, ERVK-21, ERVH-4, CYB5R2, NLK, STK26, GOLM1, SLC15A3, LINC00645, LRP1B, GALNT7, MAP3K20, KCNK9, TRG-AS1, MIR4429, ACSL5, LINC01132, ADAMTS9-AS2, RPL17-C18orf32, ERAP1, SLC25A37, ERVK-20, CDH5, CD79B, MIR1270, MIR1287, SULF2, PI4K2A, MS4A1, PSAT1, CDS1, BTBD2, MIR297, C20orf181, CERNA3, PERCC1, TPTEP2-CSNK1E, ZCCHC8, CDCA7L, ASIC4, CD22, TAX1BP3, SIK1B, G0S2, MIR6798, CD28, TMEM176A, MIR543, MIR885, NSUN5, HILPDA, FOXD1-AS1, RCC2, LANCL2, CMAS, MIR4516, KMT2E, ACSS2, CD14, MIR374B, CD19, CERS2, RIOK2, ERVK-19, TDP1, AP5M1, HHAT, MIR708, WDR11, CDO1, UBE2R2-AS1, PI15, HJURP, AGGF1, ANO1, RTN4R, STMN3, MIR513C, IFT57, FEZF2, EPN3, CD38, CDKN2C, CDKN1C, TUG1, TESC, ULK4, MIR1468, FOCAD, COMETT, APOBEC3A_B, MIR1288, HEATR1, MTPAP, F11R, MIR1224, LAPTM4B, SLC39A9, KIR2DS2, DRAM1, ACACB, ZNF654, CD24, ADI1, HMMR-AS1, ASAP1, TRAP, STEAP3, SAMMSON, IL20, MIR1225, CD33, APPL2, MFT2, MIR564, ATP1A3, MIR562, MIR215, AMOT, ABCA13, APOA1, CREBRF, APLP2, MIR211, MIR212, PPP1R1C, SIK1, MIR214, MIR217, BCR, DIRAS1, APCDD1, IL34, MIR218-2, MIR22, MIR25, CMTM2, CMTM4, ANXA1, OPN1SW, FBXO16, LINC00599, SLC7A13, PRUNE2, APOD, MIR198, PDE12, APOBEC3A, CRTC2, TXLNA, MIR199A1, ANO6, XAGE3, ZNF596, RNF168, MIR199A2, OXER1, CITED4, MIR19A, AQP8, SLFN5, MIR200B, APOB, APOA4, MIR206, ANCR, AMH, CAPNS1, CCL3L3, UHMK1, PLK5, CPT1C, MSI2, CMTM3, NAA30, LGMNP1, TNFSF12-TNFSF13, ALKBH2, H4-16, EIF2AK1, PIWIL4, EIF2AK4, ANTXR2, MRGPRX4, MRGPRX3, MIAT, BMPR1B, BATF2, CTHRC1, TRMT61A, TSGA13, PIFO, TMEM18, FBLN7, NEU4, VTI1A, ROMO1, CTCFL, GATA5, MIR301A, MIR30C1, UPRT, TMEM71, BDKRB1, BMP1, MIR30C2, TAAR1, GPR151, MIR30E, EGFLAM, AMELX, MIR320A, IL31RA, AKR1B1, MIR93, RBM45, TIGIT, CMYA5, MIR193A, SOX2-OT, LHFPL3, LINC01116, ASPG, ATF1, MIR140, ASPA, ART1, FOXD4L4, NUP43, ANKDD1A, H3P44, FOLH1B, BAAT, BACH1, ZFP57, ARNTL, ARHGAP1, RSPO2, TSPAN33, STING1, BAD, HSR, LINC-PINT, ATF3, ATF4, CEP85L, ATP1A1, ATP2A2, ATP5F1A, ALDH7A1, MIR128-1, MIRLET7G, MIRLET7E, MIR129-1, MIRLET7C, MIR129-2, MIR130A, ERVFRD-1, ATP5F1B, SOHLH1, LINC01446, BCAR4, C1QTNF8, TNFAIP8L3, C12orf75, ATP7A, MIR130B, SNAI3, ZAR1, CELIAC2, MIR15B, MCOLN2, IGFBP7-AS1, ZDHHC23, ARG1, MIR181A2, CACNA1G-AS1, RALGAPA1, EBF3, RICTOR, SEMA3D, SLC35F1, GPRC6A, ARF4, ARF1, BCAT2, KDM1B, HEPACAM, SKA1, BCL2A1, MIR184, MIR185, GK5, ANKS4B, TRIM59, NCR3, TUSC1, RHOC, SNHG15, HOXA-AS2, BAG1, TUSC7, STPG4, MIF-AS1, LINC00663, MIR7-3HG, EMC10, CXCL17, ADGRB3, NEK8, MIR150, LOC283731, RHOB, LINC01579, H19, BAK1, MRGPRX1, AQP7P3, VN1R17P, C1QTNF1, MIR505, ZNF436, CAD, TTYH3, MIR519D, CALCA, MIR517C, MIR500A, DNAJC5, ULBP1, ADCYAP1, MIR506, ADD3, HAS2-AS1, ERVK-8, TRIM45, ORAI2, PANK2, MIR455, NAA25, DHDDS, LEF1-AS1, ADAR, MIR524, HM13, SLC38A1, VOPP1, SOX7, CA2, DOHH, MXD3, EMC6, MED25, SPRY4, MIR432, VANGL1, MIR495, NUAK2, MAP1LC3B, DIAPH3, DEFB126, KAZALD1, MIR193B, MIR520A, NECTIN4, MIR520B, MIR518B, CACNA1F, BAALC, MIR125B2, NHEJ1, BRCC3, MMP28, MAPKAP1, TRIM48, CHAC1, LINC00273, CARD14, SLC25A23, CANX, ATG9A, ADA, CAPN1, AHNAK, SNHG20, GGCT, DUSP26, MIR542, HNRNPA1P10, WNK2, FRTS1, SNORD76, MIR559, FA2H, NORAD, CRNDE, ADAM8, SLTM, CARF, WIPF3, LINC02604, ZFHX4, CAMK2A, RIOX1, RASSF10, CENPU, RTL10, VTCN1, DYNC2H1, GALNT14, RNASEH2B, ADIPOR2, TCTN1, CAMK2B, CAMK2G, ULBP3, NOX5, BHLHE41, ZNHIT2, FERMT3, ELFN2, MIR373, NLRP12, SLFN11, PLXNA4, DUOXA1, MIR342, BTF3, MIR346, MIR369, AGRP, GFM1, DOCK7, CRISPLD2, RHPN2, NKD2, NKD1, ERVK-7, MIR375, ALG2, H2BC12, SNHG7, AJUBA, BTG1, NEK9, DST, SCGB3A1, BNIP3L, CYGB, MIR155HG, GPR166P, TIRAP, NLRP3, TRIM9, MIR302B, MIR302D, LINC00313, CMTM1, BMX, SLC46A1, MTFR2, CYTOR, MIR323A, FOXP2, IGSF8, NAPRT, AGXT, MARCHF9, SLC38A5, RSPO3, MIR379, PHF5A, ING5, MIR433, HSDL2, C9, ZDHHC18, RHBDD1, MAF1, USP48, PPP1R1B, KAT8, EVA1A, KBTBD7, B3GNT5, CCDC8, GRK3, KLF16, USP26, ITCH, MIR490, MIR491, RIOK1, TLCD3B, MIR363, H4C15, FOXD2-AS1, PHF6, PCGF1, MIR381, MIR422A, MIR423, CNDP1, CLEC18B, LNX1, C4BPA, KDM2B, GLIS2, ADGRE3, MIR429, MAP1LC3A, MAK16, MIR448, HOPX, MIR449A, PHYHIPL, MAP3K21, AKT1S1, BRMS1L, LINC00115, LAMP1, B3GAT1, GNB3, GPC1, GPD1, SALL2, GPM6B, CCR10, S100A8, S100A6, SORT1, GPR26, RTKN, GRK5, RRBP1, RRAS, MKNK2, RPS20, RPS15, RPS11, RPS6KA2, RPS6KA1, GPS2, RPS3, RPL17, RPL8, RPL5, RPA1, GPX3, GRIA4, SCNN1A, SCO1, SRL, SFRP2, SIM2, ST8SIA1, ST6GAL1, SIAH2, SHH, SH3GL3, GNRH1, ITSN1, SGTA, SCG5, SRSF3, SRSF1, SFRP1, GOT2, SETMAR, SEL1L, GOLGA4, SDCBP, CX3CL1, CXCL6, CCL22, CCL21, CCL20, CCL18, CCL13, GOT1, GRM1, GRM2, RGR, PTGDS, GZMM, PTPRC, PTPN14, PTPN6, PTPN3, PTPN1, CFH, HFE, PTHLH, HGFAC, NRG1, PTGFRN, TAS2R38, GZMB, PSMD12, MNX1, PSMD2, PSMC6, PSMB9, PSMB4, PSG2, PSD, PSAP, RELN, HLA-DQA1, PRPS1, PTPRF, PTPRZ2, REV3L, RARRES2, GRM3, DPF2, UPF1, REN, RELB, REG1A, RDX, RBM3, RBBP8, RBBP5, RBBP4, RASA1, RARRES1, GTF2H4, RARG, RARA, RAP1A, RANBP2, RAN, RAD51B, GSTM1, RAC2, RAB27B, RAB3A, PXN, PVT1, SIX3, GNAS, CLCN3, SLAMF1, FGF7, FGF9, TMSB4X, TMPO, TSPAN8, TLR2, TLE1, TK1, FGFR4, FHL1, THY1, THOP1, TGFB3, FOXD1, FOXC2, FLNB, TFDP1, TFAP2A, FLNC, TERF2, FLT3, TEAD4, TCP1, TCN2, FOLR1, TCF3, TCF4, TNFRSF1B, TNR, FGF5, TRPC6, TYRP1, TYROBP, TYK2, FBP1, TNFSF4, FCGR2A, TUB, FCGRT, TTC4, PHLDA2, TSC2, FDFT1, TRP-AGG2-6, FGF1, TRIO, TRH, FDXR, TRAF1, HSP90B1, TPT1, TPR, TPI1, TPD52L2, FECH, FEN1, FER, TBX2, TAT, TAP1, SLC18A1, SNAPC2, GHRHR, SMO, GJA3, GCLC, GCLM, SMARCA4, SMARCA2, SLN, SLC22A5, GLO1, SLC20A1, SLC12A4, SNRPN, SLC12A3, GLUD2, SLC7A4, SLC8A2, SLC6A8, SLC5A2, SLC4A1, SLC3A1, GML, SLC1A5, SLC1A3, SLC1A1, SNRPB, GGT7, FTH1, GABPB1, TACC1, FTL, GAST, SYK, ABCC8, STX1A, STK11, FYN, FZD2, STAT2, ST14, ST2, SST, GDF2, ITPRID2, SREBF2, GABRA1, GALR1, SPTA1, SPOCK1, GATA2, SPAG4, SP100, SOX5, SOX1, GCH1, HLA-DQB1, PROP1, PRODH, HLA-DRA, MUC4, MTRR, RNR2, IRF1, MT3, MT1E, MT1A, MSX1, MST1R, IRF5, MSR1, IRF7, MSH4, ISG20, ITGA2B, ITGA3, MPST, MPP1, MPO, MPG, MMP19, ITGA4, MMP13, MMP11, ITGA9, MMP7, ITGAE, MX2, EIF3E, MYB, NEK2, NME3, NME2, NME1, NINJ1, NGFR, IL1RN, NFKB2, NFIB, IL11, IL13RA1, NEUROD2, NELL2, IL15, MYBL2, IMPDH2, SEPTIN2, NDN, NCK1, INPP5B, NAB2, MYO1C, MYH2, MYD88, INSM1, INSR, MYBPH, ITGB1, MME, AFDN, LMNA, KIF5B, SMAD1, MXD1, KIF5C, TACSTD2, LTBR, LSS, LRP6, KLRC3, KNG1, LNPEP, LMO2, LIMK2, SMAD7, KIF11, LIFR, LGALS9, LGALS8, LGALS3BP, LAMA2, LAMA3, LAMA5, LCP2, LCN1, LASP1, STMN1, KCNMA1, MAGEA1, KMT2A, MAP3K1, MAP3K11, ITGB3, MIP, CXCL9, ITGB5, CD99, ITGB8, KITLG, MGAT5, MGAT1, ITK, MEOX2, ANOS1, MAGEA2, KCNA5, KCNH1, MCM6, MAZ, MAX, MATK, MAP2, MAP1B, KCNJ3, MAOA, MAN2A1, MAL, NOP2, NOS3, IGSF1, PLD2, POU1F1, PON2, POLR2F, HOXA13, SEPTIN4, HOXB@, HOXB1, PNOC, PLXNA2, PLP2, PLOD2, PLEC, PLCG1, HOXA1, HOXB4, HOXB5, PLA2G4A, HOXB6, PKD2, PITX1, PIP4K2A, PIK3R2, HOXB9, HOXC9, HOXC11, PIK3C3, POU2F1, HNRNPU, SERPINE2, PRKCE, PRLR, PRL, EIF2AK2, MAP2K3, HLA-E, MAPK11, HLA-F, HMGB2, HMGB3, PRKCZ, HMOX2, PRKCG, PRKCD, PPIB, NR4A1, FOXA1, HNF4A, HNRNPA1, HNRNPC, SRGN, PRELP, PPT1, PPP2R5E, HNRNPL, PPP1CA, PPID, SERPINI1, SERPINB6, NPY, NR4A2, PAM, IARS1, PAFAH1B1, PCSK6, P2RX7, OXTR, OPRM1, OPRL1, OPCML, OLR1, OCRL, OCA2, IFN1@, REG3A, NUCB2, IFNA17, IGFALS, ROR2, IGFBP6, NPTX2, NPPA, NPAS2, PNP, NOVA1, CCN3, IGHA1, CNTN3, NDST1, PI3, PRMT2, SLC25A3, HOXD9, HOXD10, PGAM1, PFN1, PFKFB4, HPCAL1, PFKFB2, SERPINF1, PDK4, PDK3, HRG, HSPA8, PAX3, PDE4D, PDE4A, HSPA9, PCYT1A, CDK18, HSP90AB1, PCDH8, PCBP2, PBX3, PDCL, PAX7, HSPG2, UBE2I, UBE2N, UGDH, HDAC5, MAP3K2, CTNND1, TCFL5, OGA, CCT8, TNFSF13B, CD226, CTCF, CTSG, CELF2, CTSS, CUX1, IGF2BP3, RAD51AP1, RASL10A, CYB561, CYBB, TXNIP, CYC1, MXD4, CYLD, SORBS1, CCT2, CCT7, CYP1B1, DPYSL4, ARFGEF1, CHL1, TRAF3IP2, CTNNA1, CTAA1, DEPP1, UBE2C, CSNK1E, ADRM1, CSNK2A1, KDELR2, TLK2, CSNK2A2, CST3, SLC38A3, COPS6, RAB40B, PAPOLA, ZMYND11, BRD8, MMP24, PPARGC1A, MRPS30, SMR3B, CTBP1, CYP46A1, CTH, LAMC1, TUBGCP2, GJB6, SEPTIN9, ARFGEF2, PROCR, ANP32B, TRIM28, RAMP3, ABCC4, SPRY2, SPRY1, SPRY3, MSLN, DBH, MPZL2, CALCRL, TSHZ1, RNF41, ATP6AP2, DBN1, SEMA3A, PDIA6, ARL4C, PPIF, ACTR3, PTPRU, DNM1L, DBT, ABCC5, PARP3, CHAF1A, FRAT1, PDCD6, TFG, DAG1, RNASEH2A, CYP4B1, CYP2A7, HYOU1, KAT5, CYP2B6, CYP2E1, SEMA6B, CREB3, CAP1, HOXB13, EIF3M, ZBTB18, FST, GPNMB, TUBB4A, HAX1, LRRN2, MCRS1, C1D, RBM14, TUBGCP3, CYP27A1, PIAS3, RACK1, MYL9, DAB1, DLC1, TRIOBP, CD160, KIF3A, ATP6V0A2, CADM1, RAB38, SMPX, CCR1, PPP1R15A, LY96, DAPK2, CCR6, DDX58, LTB4R, LPAR3, WBP2, MAPK8IP2, KIF4A, SEC14L2, SRRM2, TNFRSF13B, SEC61G, COL4A1, TRAM1, COL4A3, COL16A1, ANGPTL2, HARS2, TARDBP, RYBP, BCL2L13, RASGRP3, COX5B, CLK2, NOX1, PABPC1, HBP1, RANBP6, SNORD48, INTS6, GNL3, HSPB8, DGCR5, NOC2L, PHF19, ZBTB20, RAI14, HECTD1, DNM3, CHD5, TPP1, SNED1, ULK3, ADGRA2, RWDD3, SYF2, MYRIP, CLTC, MPC2, PRKD2, COL17A1, COX7A2, ZWINT, CBX3, CRH, TPX2, CCT5, SIRT2, ATF6, CD93, CRKL, CRMP1, ZHX2, CILK1, CHSY1, CRYZ, PDAP1, DAAM1, PARK7, RNF139, CS, POLG2, FZD10, CSE1L, CSF1, PTENP1, RASSF1, TREH, PSIP1, CSHL1, KDM2A, CNOT1, SIRT5, SULF1, CP, TNS2, PUM2, PPP1R13B, PSD3, HAUS5, DOCK9, SYNE1, SYNM, ATMIN, CAMTA1, PLCB1, JMJD6, CPT1A, ATG4B, CPB2, LARP4B, DIP2A, CLDN4, SSPO, MAST2, CPS1, ARHGAP26, SWAP70, SMG1, WDTC1, PDCD6IP, DCK, UGP2, PTBP3, ITGA10, PARG, CDC42BPA, FOXN1, CUL1, CUL3, ELAVL1, NR0B2, SPARCL1, MAD1L1, H4C14, H4C13, H4C5, H4C2, H4C8, H4C3, H4C11, H4C12, H4C6, H4C4, H4C1, ELK4, ELN, ENDOG, ENO1, ENO2, ENPEP, DGKZ, API5, SERPINB1, EFNB2, FADD, RIPK2, CD164, ADAM23, EEF2, TNFSF12, TNFSF14, HRK, EFNA2, RNMT, ADAM19, EFNB1, VAMP8, CDC14B, EIF3A, IRS2, TNKS, EIF4G2, DCHS1, STC2, PLPP3, PLPP1, CASK, KHSRP, DENR, DEGS1, FZD4, CDC7, H4C9, VIPR1, ETV2, ETV4, WNT10B, WNT7B, WNT6, EXTL3, WNT2, F2, F2RL1, NAT2, VRK1, F9, VGF, YES1, VEGFB, FABP4, FABP5, VCL, FANCA, KDM6A, UTRN, FAH, USF1, UROS, USP4, UNG, XPC, YWHAG, TKTL1, ERG, SMC1A, EP300, ESS2, BAS, EPHA7, DPF1, AKAP1, EPHA8, ERCC2, CCDC6, KAT6A, EREG, ESD, YWHAZ, DEK, USP7, ETFA, BTG2, LAPTM5, PRDM2, ZYX, ZNF217, ZBTB16, ZNF131, ZNF22, CNBP, DLK1, EEF1B2P2, SUCLG1, EIF4E2, KLK4, NCOR2, DHPS, TP53I3, POLR1C, DIAPH1, TMEM59, DIO1, ADAMTS1, ADAMTS4, CABP1, DIO2, PICK1, MICAL2, EIF2AK3, MAP4K4, DLG3, QKI, NCR1, NCR2, DLX2, SARDH, FADS2, DYNC1I1, RECQL4, SLC22A8, DFFB, EIF5B, TGM5, ARHGEF11, HNRNPDL, NR1D2, ACE, USP3, KIF14, ARNT2, UBAP2L, SNAP91, DDB2, CEP350, GAB2, TRIM14, SPATA2, PHF14, RB1CC1, MTSS1, DLGAP5, EIF4A3, PCLAF, DDX6, DMXL1, DOCK4, SEMA3E, ULK2, ST18, ESPL1, ADIPOQ, DPP4, GALR2, E2F6, CCRL2, PKD2L1, CH25H, DVL2, TRPA1, DVL3, PLOD3, USP13, BTRC, KYNU, E2F2, VNN2, SYNJ2, SPAG9, IER3, SYNJ1, PER3, ECE1, ASAP2, ECHS1, ALKBH1, CCN4, GGH, EDN3, NRP2, CDKL1, UBA3, PRC1, NREP, DGKI, SOCS6, GPR55, GPR37L1, PIWIL1, STK17A, MAPKAPK2, SLC26A3, NOG, IL32, DLGAP1, HACD1, RCAN1, EXO1, SLC7A7, CBFA2T2, AIFM1, PDLIM1, DUSP4, NMI, FCGR2C, USP8, USP10, SART1, PKMYT1, DIRAS3, SYT7, VSNL1
-
Diploid Triploid Mosaic
Wikipedia
See also: Triploid syndrome This article is an orphan , as no other articles link to it . ... Individuals with diploid-triploid syndrome have some cells with three copies of each chromosome for a total of 69 chromosomes (called triploid cells) and some cells with the usual 2 copies of each chromosome for a total of 46 chromosomes (called diploid cells). [1] Having two or more different cell types is called mosaicism . ... A well-known example of trisomy is trisomy 21 or Down syndrome . [2] References [ edit ] ^ a b "Diploid-triploid mosaicism" .
-
Myelokathexis
Wikipedia
The disorder shows prominent neutrophil morphologic abnormalities. [ citation needed ] Myelokathexis is amongst the diseases treated with bone marrow transplantation and cord blood stem cells . [ citation needed ] WHIM syndrome is a very rare variant of severe congenital neutropenia that presents with warts, hypogammaglobunemia, infections, and myelokathexis. ... The truncated form of the receptor has a 2-fold increase in G-protein coupled intracellular signalling, and this mutation of the receptor can be identified by DNA sequencing. [3] See also [ edit ] WHIM syndrome References [ edit ] ^ a b Aprikyan AA, Liles WC, Park JR, Jonas M, Chi EY, Dale DC (Jan 2000). ... "Mutations in the chemokine receptor gene CXCR4 are associated with WHIM syndrome, a combined immunodeficiency disease".
-
Fibrous Dysplasia/mccune-Albright Syndrome
Gene_reviews
Recommended evaluations for growth hormone excess in individuals with fibrous dysplasia/McCune-Albright syndrome Figure 6. Recommended evaluations for gonadal abnormalities in females with fibrous dysplasia/McCune-Albright syndrome Figure 7. Recommended evaluations for gonadal abnormalities in males with fibrous dysplasia/McCune-Albright syndrome Figure 8. Recommended evaluations for thyroid abnormalities in individuals with fibrous dysplasia/McCune-Albright syndrome Figure 9. ... Recommended management for fibrous dysplasia in individuals with fibrous dysplasia/McCune-Albright syndrome Endocrinopathies Precocious puberty. ... Recommended management for precocious puberty in girls with fibrous dysplasia/McCune-Albright syndrome Figure 13. Recommended management for gonadal involvement in boys with fibrous dysplasia/McCune-Albright syndrome Thyroid disease. ... Recommended management for growth hormone excess in individuals with fibrous dysplasia/McCune-Albright syndrome FGF23-mediated phosphate wasting.
-
Transient Global Amnesia
Wikipedia
The leading hypotheses are some form of epileptic event, a problem with blood circulation around, to or from the brain, or some kind of migraine-like phenomenon. [7] [14] [15] [16] The differences are sufficiently meaningful that transient amnesia may be considered a heterogeneous clinical syndrome [2] with multiple etiologies, corresponding mechanisms, and differing prognoses. [8] Precipitating events [ edit ] TGA attacks are associated with some form of precipitating event in at least one-third of cases. [17] The most commonly cited precipitating events include vigorous exercise (including sexual intercourse), swimming in cold water or enduring other temperature changes, and emotionally traumatic or stressful events. [2] There are reports of TGA-like conditions following certain medical procedures and disease states. [15] One study reports two cases of familial incidence (in which two members of the same family experienced TGA), out of 114 cases considered. [2] This indicates the possibility that there could be a slight familial incidence. ... Diagnosis [ edit ] Differential diagnosis [ edit ] A differential diagnosis should include: [29] Thrombosis of the basilar artery Cardioembolic stroke Complex partial seizures Frontal lobe epilepsy Lacunar syndromes Migraine variants Posterior cerebral artery stroke Syncope and related paroxysmal spells Temporal lobe epilepsy If the event lasts less than one hour, transient epileptic amnesia (TEA) might be implicated. [2] [30] If the condition lasts longer than 24 hours, it is not considered TGA by definition. ... PMID 6658982 . ^ a b c d e f g h i j k l m n o p Hodges; Warlow, CP (1990). "Syndromes of transient amnesia: towards a classification. ... External links [ edit ] Classification D ICD - 10 : G45.4 ICD - 9-CM : 437.7 MeSH : D020236 DiseasesDB : 13251 External resources eMedicine : neuro/380 v t e Cerebrovascular diseases including stroke Ischaemic stroke Brain Anterior cerebral artery syndrome Middle cerebral artery syndrome Posterior cerebral artery syndrome Amaurosis fugax Moyamoya disease Dejerine–Roussy syndrome Watershed stroke Lacunar stroke Brain stem Brainstem stroke syndrome Medulla Medial medullary syndrome Lateral medullary syndrome Pons Medial pontine syndrome / Foville's Lateral pontine syndrome / Millard-Gubler Midbrain Weber's syndrome Benedikt syndrome Claude's syndrome Cerebellum Cerebellar stroke syndrome Extracranial arteries Carotid artery stenosis precerebral Anterior spinal artery syndrome Vertebrobasilar insufficiency Subclavian steal syndrome Classification Brain ischemia Cerebral infarction Classification Transient ischemic attack Total anterior circulation infarct Partial anterior circulation infarct Other CADASIL Binswanger's disease Transient global amnesia Haemorrhagic stroke Extra-axial Epidural Subdural Subarachnoid Cerebral/Intra-axial Intraventricular Brainstem Duret haemorrhages General Intracranial hemorrhage Aneurysm Intracranial aneurysm Charcot–Bouchard aneurysm Other Cerebral vasculitis Cerebral venous sinus thrombosis v t e Human memory Basic concepts Encoding Storage Recall Attention Consolidation Neuroanatomy Types Sensory Echoic Eidetic Eyewitness Haptic Iconic Motor learning Visual Short-term " The Magical Number Seven, Plus or Minus Two " Working memory Intermediate Long-term Active recall Autobiographical Explicit Declarative Episodic Semantic Flashbulb Hyperthymesia Implicit Meaningful learning Personal-event Procedural Rote learning Selective retention Tip of the tongue Forgetting Amnesia anterograde childhood post-traumatic psychogenic retrograde transient global Decay theory Forgetting curve Interference theory Memory inhibition Motivated forgetting Repressed memory Retrieval-induced forgetting Selective amnesia Weapon focus Memory errors Confabulation False memory Hindsight bias Imagination inflation List of memory biases Memory conformity Mere-exposure effect Misattribution of memory Misinformation effect Source-monitoring error Wernicke–Korsakoff syndrome Research Art of memory Memory and aging Deese–Roediger–McDermott paradigm Exceptional memory Indirect tests of memory Lost in the mall technique Memory disorder Memory implantation Methods used to study memory The Seven Sins of Memory Effects of exercise on memory In society Collective memory Cultural memory False memory syndrome Memory and social interactions Memory sport Politics of memory Shas Pollak World Memory Championships Related topics Absent-mindedness Atkinson–Shiffrin memory model Context-dependent memory Childhood memory Cryptomnesia Effects of alcohol Emotion and memory Exosomatic memory Flashbacks Free recall Involuntary memory Levels-of-processing effect Memory and trauma Memory improvement Metamemory Mnemonic Muscle memory Priming Intertrial Prospective memory Recovered-memory therapy Retrospective memory Sleep and memory State-dependent memory Transactive memory People Robert A.
-
Mosquito Bite Allergy
Wikipedia
The development of desensitization that follows repetitive mosquito bites and reduces the intensity or completely blocks reactions to mosquito bites may take longer to develop and/or be less effective in those with Skeeter syndrome compared to those with ordinary mosquito bite reactions. [11] Diagnosis [ edit ] The diagnosis of Skeeter syndrome is based mainly on the appropriate history of severe skin responses to mosquito bites that may be associated with fever. ... PMID 25758116 . ^ Simons FE, Peng Z (September 1999). "Skeeter syndrome". The Journal of Allergy and Clinical Immunology . 104 (3 Pt 1): 705–7. doi : 10.1016/S0091-6749(99)70348-9 . ... PMID 24662020 . ^ a b c d e f g h i j k Weins AB, Biedermann T, Weiss T, Weiss JM (October 2016). "Wells syndrome" . Journal der Deutschen Dermatologischen Gesellschaft . 14 (10): 989–993. doi : 10.1111/ddg.13132 . ... "Treatment of eosinophilic cellulitis (Wells syndrome) - a systematic review". Journal of the European Academy of Dermatology and Venereology : JEADV . 30 (9): 1465–79. doi : 10.1111/jdv.13706 . ... "Cellulitis, Eosinophilic (Wells syndrome)". StatPearls . PMID 30335327 .
-
Secondary Hypertension
Wikipedia
People with neurogenic hypertension respond poorly to treatment with diuretics as the underlying cause of their hypertension is not addressed. [13] Pheochromocytoma – a tumor which results in an excessive secretion of norepinephrine and epinephrine which promotes vasoconstriction Hyperaldosteronism ( Conn's syndrome ) – idiopathic hyperaldosteronism, liddle's syndrome (also called pseudoaldosteronism), glucocorticoid remediable aldosteronism Cushing's syndrome – an excessive secretion of glucocorticoids causes the hypertension Hyperparathyroidism Acromegaly Hyperthyroidism Hypothyroidism Adrenal [ edit ] A variety of adrenal cortical abnormalities can cause hypertension, In primary aldosteronism there is a clear relationship between the aldosterone-induced sodium retention and the hypertension. [14] Congenital adrenal hyperplasia , a group of autosomal recessive disorders of the enzymes responsible for steroid hormone production, can lead to secondary hypertension by creating atypically high levels of mineralocorticoid steroid hormones. ... Cortisol is a hormone secreted by the cortex of the adrenal glands . Cushing's syndrome can be caused by taking glucocorticoid drugs, or by tumors that produce cortisol or adrenocorticotropic hormone (ACTH). [32] More than 80% of patients with Cushing's syndrome develop hypertension., [33] which is accompanied by distinct symptoms of the syndrome, such as central obesity , lipodystrophy , moon face , sweating , hirsutism and anxiety . [34] Neuroendocrine tumors are also a well known cause of secondary hypertension. ... "Renal manifestations of the antiphospholipid syndrome". Current Rheumatology Reports . 11 (1): 52–60. doi : 10.1007/s11926-009-0008-2 . ... "A Switch in the Mechanism of Hypertension in the Syndrome of Apparent Mineralocorticoid Excess" . ... "Association between Crohn's disease and Conn's syndrome. A report of two cases". Panminerva Medica . 47 (1): 61–4.
-
Alcohol Dependence
Wikipedia
Gross [19] collaborated to produce a formulation of what had previously been understood as ‘alcoholism’ – the alcohol dependence syndrome . The alcohol dependence syndrome was seen as a cluster of seven elements that concur. It was argued that not all elements may be present in every case, but the picture is sufficiently regular and coherent to permit clinical recognition. The syndrome was also considered to exist in degrees of severity rather than as a categorical absolute. ... In the DSM-5, the term addiction is synonymous with the classification of severe substance-use disorder. ^ http://www.alcoholcostcalculator.org/business/about/dsm.html ^ "ICD-9-CM Diagnosis Codes 303.* : Alcohol dependence syndrome" . www.icd9data.com . ^ Clark, David, Background Briefing, Alcohol Dependence, Drink and Drug News, 7 February 2005, p. 11 ^ "Alcohol use screening tests – GOV.UK" . www.gov.uk . ... Retrieved 16 April 2010 . ^ Edwards G. & Gross MM, Alcohol dependence: provisional description of a clinical syndrome, BMJ 1976; i: 1O58-106 ^ Alcohol Recovery Program External links [ edit ] Classification D ICD - 10 : F10 .2 ICD - 9-CM : 303 OMIM : 103780 Arnold Little, MD Alcohol Dependence – extensive article SADD – Short Alcohol Dependence Data Questionnaire . ... ISBN 5-222-04382-7 v t e Psychoactive substance-related disorder General SID Substance intoxication / Drug overdose Substance-induced psychosis Withdrawal : Craving Neonatal withdrawal Post-acute-withdrawal syndrome (PAWS) SUD Substance abuse / Substance-related disorders Physical dependence / Psychological dependence / Substance dependence Combined substance use SUD Polysubstance dependence SID Combined drug intoxication (CDI) Alcohol SID Cardiovascular diseases Alcoholic cardiomyopathy Alcohol flush reaction (AFR) Gastrointestinal diseases Alcoholic liver disease (ALD): Alcoholic hepatitis Auto-brewery syndrome (ABS) Endocrine diseases Alcoholic ketoacidosis (AKA) Nervous system diseases Alcohol-related dementia (ARD) Alcohol intoxication Hangover Neurological disorders Alcoholic hallucinosis Alcoholic polyneuropathy Alcohol-related brain damage Alcohol withdrawal syndrome (AWS): Alcoholic hallucinosis Delirium tremens (DTs) Fetal alcohol spectrum disorder (FASD) Fetal alcohol syndrome (FAS) Korsakoff syndrome Positional alcohol nystagmus (PAN) Wernicke–Korsakoff syndrome (WKS, Korsakoff psychosis) Wernicke encephalopathy (WE) Respiratory tract diseases Alcohol-induced respiratory reactions Alcoholic lung disease SUD Alcoholism (alcohol use disorder (AUD)) Binge drinking Caffeine SID Caffeine-induced anxiety disorder Caffeine-induced sleep disorder Caffeinism SUD Caffeine dependence Cannabis SID Cannabis arteritis Cannabinoid hyperemesis syndrome (CHS) SUD Amotivational syndrome Cannabis use disorder (CUD) Synthetic cannabinoid use disorder Cocaine SID Cocaine intoxication Prenatal cocaine exposure (PCE) SUD Cocaine dependence Hallucinogen SID Acute intoxication from hallucinogens (bad trip) Hallucinogen persisting perception disorder (HPPD) Nicotine SID Nicotine poisoning Nicotine withdrawal SUD Nicotine dependence Opioids SID Opioid overdose SUD Opioid use disorder (OUD) Sedative / hypnotic SID Kindling (sedative–hypnotic withdrawal) benzodiazepine : SID Benzodiazepine overdose Benzodiazepine withdrawal SUD Benzodiazepine use disorder (BUD) Benzodiazepine dependence barbiturate : SID Barbiturate overdose SUD Barbiturate dependence Stimulants SID Stimulant psychosis amphetamine : SUD Amphetamine dependence Volatile solvent SID Sudden sniffing death syndrome (SSDS) Toluene toxicity SUD Inhalant abuse v t e Reinforcement disorders: Addiction and Dependence Addiction Drug Alcohol Amphetamine Cocaine Methamphetamine Methylphenidate Nicotine Opioid Behavioral Financial Gambling Shopping Palatable food Sex-related Intercourse Pornography Internet-related Internet addiction disorder Internet sex addiction Video game addiction Digital media addictions Cellular mechanisms Transcriptional ΔFosB c-Fos Cdk5 CREB GluR2 NF-κB Epigenetic G9a G9a-like protein HDAC1 HDAC2 HDAC3 HDAC4 HDAC5 HDAC9 HDAC10 SIRT1 SIRT2 ...GABRA2, ALDH2, HTR2A, ADH1C, ADH1B, CYP2E1, OPRM1, NPY, PDYN, SLC6A4, SNCA, CHRNA5, GABBR1, TACR1, TAS2R38, CCKAR, CHRNA3, NPY2R, GABRG2, SLC29A1, SHBG, TACR3, GGT1, ADH4, FTO, SERINC2, CTNNA2, KIAA0040, PKNOX2, LINC02694, KCNJ6, THSD7B, AKR1A1, BDNF, CHRM2, POMC, DBH, MAOA, ANKK1, HTR1B, DRD2, DRD3, DRD4, CRHR1, COMT, SLC6A3, OPRK1, CRH, ALDH1A1, MAOB, GRIN2B, CNR1, TPH1, ADH7, HTR2C, TH, HTR1A, MTHFR, NFKB1, TRH, GATA4, GAD1, APOE, GRIN2A, OPRL1, GABRB1, IL6, GABRA6, HTR3A, CCK, MPDZ, GABRB3, GABRA1, HTR7, SGIP1, IL1RN, IL10, GRM8, LEP, IL1B, GRIN1, MMP9, OPRD1, NR4A2, GLUL, GH1, NTRK2, GAL, GAD2, GABRG1, DRD1, HNMT, SLC6A2, ADH1A, CLOCK, HTR3B, CHRNB4, ADH5, ARSA, CHRNA4, ZNF699, SLC18A2, GRIK1, SNRNP70, GABRB2, SRD5A1, TAC1, CDH11, CDH13, GABRG3, ACE, GRIK3, SLC1A2, TP53, GRM1, TTC12, PTP4A1, ADRA2A, IL1R1, IL1A, SGCE, AKR1C3, SDHAF3, ABO, HMGB1, TKT, CAT, FYN, NTSR1, PHF3, CNTNAP2, DKK2, OXT, CREB1, CHRNB3, CRHBP, NTS, GHS, SLC6A5, RFX4, PENK, XRCC5, KPNA3, AGO1, LRP8, UBAP2, SEMA5A, CXCL8, SLC17A5, GEMIN4, TESK2, TIPARP, PIK3R1, SAT1, LILRA1, KLF11, GALR3, MGLL, GALR2, ZCCHC14, ANKRD7, ARC, RGS4, NRXN3, SLCO3A1, NCAM1, NQO2, TAS2R16, SIGMAR1, KANK1, SLC6A9, C1D, IPO11, SRD5A2, HERPUD1, PCDH12, NEUROD2, PDE10A, AGO2, TFAP2B, SPG21, CYTL1, CARTPT, NRDC, MOG, MOBP, HOMER1, SLC6A1, NPY5R, CNTN6, NAT1, DSCAML1, EPHX1, GRM3, GABRR1, GRM2, CAMK2A, NKAIN1, THEMIS, DPYSL2, OSBPL5, CYP2A13, CDH12, CDH15, GNB3, CNTN4, GLI2, GHSR, GABRA5, AKR1C4, CNR2, NKAIN2, GAPDH, GAP43, GALR1, CALCA, CAMK4, DUSP8, PTK2B, HLA-DRA, PER3, ADCY7, ADH6, NLGN4X, STON2, HAMP, ALDH3B2, ALK, GSTM1, DTNBP1, AR, GRM7, FABP2, CASC4, ASTN1, GABRR2, EP300, EGF, RFC1, CYP2B6, SLC46A1, EGFR, RASGRF2, ECHS1, PDE4B, REN, CDK20, RACK1, CFTR, ADCY5, TBX19, VWF, PHLDA2, SNORA54, NPS, BAG3, MIR382, BHMT, TF, ST18, CARS1, GPHN, NPSR1, CDH5, CDH8, CDH9, EPHA8, CDH18, CDH10, GGH, FOLR1, MMP2, MBP, NAP1L4, FKBP5, LHB, GFAP, IL17A, FSHB, ANAPC1, TAGLN3, PCDH10, PPP1R1B, HDAC2, NMUR2, SLC22A18, AVPR1B, BRAP, SEMA3A, UTP20, ARL15, AGBL4, STAT3, RARA, PECR, LHPP, MREG, ANKS1B, KLF12, PML, STK40, C1orf220, CCSER1, FIP1L1, NCALD, FSTL5, AVP, NUMA1, NRXN1, PPP1R16B, RHOG, SLC39A8, GSS, STX18-AS1, TRPC4AP, LINC02268, ZBTB16, FAM162A, LINC01818, LINC02661, ESRRG, RN7SL697P, ADAMTSL1, AOX3P, PLGRKT, NSG1, AOX3P-AOX2P, STAT5B, BCOR, MBNL2, SLC6A6, C15orf32, NPM1, GCKR, STAG3, CSRNP3, IGSF22, IGSF9B, PRKAR1A, IRF2BP2, SETD5, TBL1XR1, GRK5, MICB, NCOA6, LYZ, MAP3K4, PLCL2, NABP1, RHBDL2, TMEM260, C16orf72, GRM5, ALLC, DDX53, LINC02210-CRHR1, LOC110806262, PRL, TSPO, SAGE1, GLP1R, GYPE, GYPB, GYPA, TPH2, FLNA, OR2AG1, FAAH, MIR21, ADIPOQ, KL, PER2, PPARA, F9, PNOC, TDO2, CCKBR, APRT, RET, TLR4, SMPD1, SMARCA1, PER1, CFP, PRDM2, NGF, CYP2A6, CCDC6, PTCH1, ESR1, DMTN, F2, EBPL, EPO, IL18R1, WDR20, ELK3, FAT1, PLCD3, EDNRB, ATN1, NLRP3, DNASE1L3, MRGPRF, DBI, OPN4, NPL, FGF2, PARP9, HTT, PCDH19, HCRTR1, CPNE5, HARS1, GUSB, GSTT1, GSR, DCLRE1C, GSK3B, NR3C1, GRIN2C, EFHD2, GPT, GM2A, GDNF, OPA3, EFHC2, PNPLA3, SLC19A3, GCG, CYP3A5, FN1, CYP19A1, COL6A3, CNIH3, AGT, ARNTL, LINC00273, GGTLC5P, GGTLC3, GGT2, GGTLC4P, AIRE, MIR4456, ALDH1B1, THRA1/BTR, MDD2, AGER, AP2B1, ADCYAP1, RN7SL263P, ADCY9, ADCY1, ADA, STIN2-VNTR, LOC111216288, OPN1SW, MDD1, TMEM161B, CDK5, CRP, ZNF366, HHEX, CHRNB1, H19, CHRM5, BTBD8, EYS, CHM, CD40, DST, CD36, CACNA1C, DAGLA, BRCA1, MIR126, MIR141, MIR155, MIR183, MIR19A, NLN, RETN, SLC17A6, PTPN11, RXRB, ARFGEF2, PDLIM5, RNU1-4, BRD2, PPARGC1A, PRSS21, RAB40B, SPACA9, SCN11A, SRSF5, MAPK8, PPAT, KDM6B, PPARG, PPARD, ABAT, PLG, HEY2, RBFOX2, SORT1, SLC1A3, TFIP11, TIMP1, NOL3, ST8SIA4, HGS, XRCC4, UMOD, DLGAP2, PPIG, TYR, TNF, THRA, SMS, THOP1, TGFB1, TFF3, TAT, SYN2, HDAC6, EBI3, SST, DHRS9, PIK3CG, PIK3CD, HLA-B, IL16, MFAP1, MEF2C, MC4R, MARK1, LOX, ABLIM1, KCNN3, KCNK3, IMPA1, IL12B, GDAP1, HTR1E, ACSS2, HSPG2, NPDC1, KCNK13, ARHGEF7, HSD11B2, HRAS, HP, MYC, MYT1, PIK3CB, HPGDS, PIK3CA, AUTS2, PHEX, PGC, PECAM1, PDGFRB, PDE4A, SALL3, PC, OXTR, NF1, NUCB2, NRGN, NPY1R, NOS3, HDGFL3, NGFR, ASCC1, HERC5, NF2, H3P40
-
Glucagonoma
Wikipedia
Typically associated with a rash called necrolytic migratory erythema , weight loss, and mild diabetes mellitus, most people with glucagonoma contract it spontaneously. [1] However, about 10% of cases are associated with multiple endocrine neoplasia type 1 (MEN-1) syndrome. [1] Contents 1 Causes 2 Mechanism 3 Diagnosis 4 Treatment 5 History 6 References 7 External links Causes [ edit ] Although the cause of glucagonoma is unknown, some genetic factors may lead to the condition. ... Glucagonoma accounts for approximately 1% of neuroendocrine tumors, although this may be an underestimate given that glucagonoma is associated with non-specific symptoms. [2] References [ edit ] ^ a b c Vinik, Aaron; Pacak, Karel; Feliberti, Eric; Perry, Roger R. (2000), Feingold, Kenneth R.; Anawalt, Bradley; Boyce, Alison; Chrousos, George (eds.), "Glucagonoma Syndrome" , Endotext , MDText.com, Inc., PMID 25905270 , retrieved 2019-09-24 ^ a b c "Glucagonoma: Practice Essentials, Pathophysiology, Epidemiology" . 2019-02-01. ... PMID 26773171 . ^ a b c d van Beek AP, de Haas ER, van Vloten WA, Lips CJ, Roijers JF, Canninga-van Dijk MR (November 2004). "The glucagonoma syndrome and necrolytic migratory erythema: a clinical review" . ... "Glucagonoma and the glucagonoma syndrome" . Oncology Letters . 15 (3): 2749–2755. doi : 10.3892/ol.2017.7703 . ... PMID 29435000 . ^ a b Adams, David R.; Miller, Jeffrey J.; Seraphin, Kathryn E. (2005-10-01). "Glucagonoma syndrome". Journal of the American Academy of Dermatology . 53 (4): 690–691. doi : 10.1016/j.jaad.2005.04.071 .
-
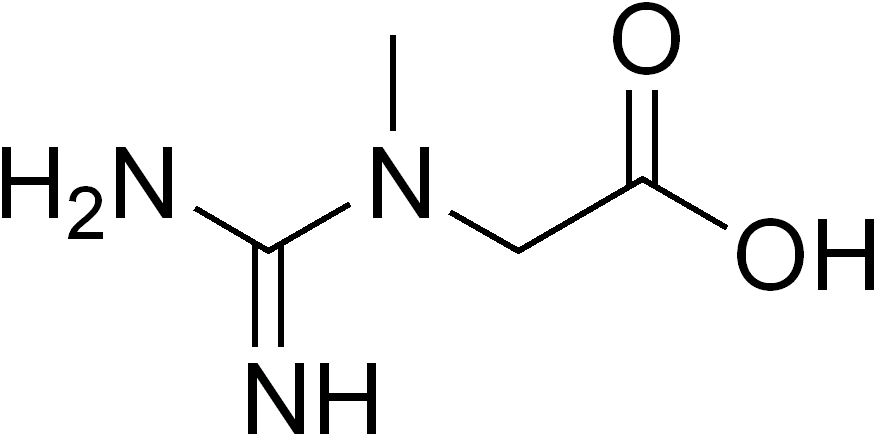
Guanidinoacetate Methyltransferase Deficiency
Wikipedia
PMID 8651275 . ^ a b c "612736 CEREBRAL CREATINE DEFICIENCY SYNDROME 2; CCDS2" . Johns Hopkins University . Retrieved 2019-01-05 . ^ a b c d Schulze, Andreas (2009). "Creatine Deficiency Syndromes". In Sarafoglou, Kiriakie; Hoffmann, Georg F.; Roth, Karl S. ... ISBN 978-0-07-143915-2 . ^ Braissant, Olivier; Henry, Hugues; Béard, Elidie; Uldry, Joséphine (2011). "Creatine deficiency syndromes and the importance of creatine synthesis in the brain" (PDF) . ... "Diagnostic methods and recommendations for the cerebral creatine deficiency syndromes". Pediatric Research . 77 (3): 398–405. doi : 10.1038/pr.2014.203 . ... Genetics Home Reference - Guanidinoacetate methyltransferase deficiency GeneReview/NIH/UW entry on Cerebral Creatine Deficiency syndromes External links [ edit ] Classification D ICD - 10 : E72.8 OMIM : 601240 MeSH : C537622 DiseasesDB : 5461 External resources GeneReviews : Creatine Deficiency Syndromes Orphanet : 382
-
Nasal Septum Deviation
Wikipedia
By itself, a deviated septum can go undetected for years and thus be without any need for correction. [4] Causes [ edit ] It is most frequently caused by impact trauma , such as by a blow to the face . [4] It can also be a congenital disorder , caused by compression of the nose during childbirth. [4] Deviated septum is associated with genetic connective tissue disorders such as Marfan syndrome , Homocystinuria and Ehlers–Danlos syndrome . [5] Diagnosis [ edit ] Nasal septum deviation is the most common cause of nasal obstruction. [6] A history of trauma to the nose is often present including trauma from the process of birth or microfractures. [6] A medical professional, such as an otorhinolaryngologist (ears, nose, and throat doctor), typically makes the diagnosis after taking a thorough history from the affected person and performing a physical examination. [6] Imaging of the nose is sometimes used to aid in making the diagnosis as well. [6] Treatment [ edit ] Medical therapy with nasal sprays including decongestants , antihistamines , or nasal corticosteroid sprays is typically tried first before considering a surgical approach to correct nasal septum deviation. [6] Medication temporarily relieves symptoms, but does not correct the underlying condition. ... "Non-cardiac manifestations of Marfan syndrome" . Annals of Cardiothoracic Surgery (Review). 6 (6): 599–609. doi : 10.21037/acs.2017.10.02 . ... External links [ edit ] Classification D ICD - 10 : J34.2 ICD - 9-CM : 470 Wikimedia Commons has media related to Nasal septum deviation . v t e Diseases of the respiratory system Upper RT (including URTIs , common cold ) Head sinuses Sinusitis nose Rhinitis Vasomotor rhinitis Atrophic rhinitis Hay fever Nasal polyp Rhinorrhea nasal septum Nasal septum deviation Nasal septum perforation Nasal septal hematoma tonsil Tonsillitis Adenoid hypertrophy Peritonsillar abscess Neck pharynx Pharyngitis Strep throat Laryngopharyngeal reflux (LPR) Retropharyngeal abscess larynx Croup Laryngomalacia Laryngeal cyst Laryngitis Laryngopharyngeal reflux (LPR) Laryngospasm vocal cords Laryngopharyngeal reflux (LPR) Vocal fold nodule Vocal fold paresis Vocal cord dysfunction epiglottis Epiglottitis trachea Tracheitis Laryngotracheal stenosis Lower RT / lung disease (including LRTIs ) Bronchial / obstructive acute Acute bronchitis chronic COPD Chronic bronchitis Acute exacerbation of COPD ) Asthma ( Status asthmaticus Aspirin-induced Exercise-induced Bronchiectasis Cystic fibrosis unspecified Bronchitis Bronchiolitis Bronchiolitis obliterans Diffuse panbronchiolitis Interstitial / restrictive ( fibrosis ) External agents/ occupational lung disease Pneumoconiosis Aluminosis Asbestosis Baritosis Bauxite fibrosis Berylliosis Caplan's syndrome Chalicosis Coalworker's pneumoconiosis Siderosis Silicosis Talcosis Byssinosis Hypersensitivity pneumonitis Bagassosis Bird fancier's lung Farmer's lung Lycoperdonosis Other ARDS Combined pulmonary fibrosis and emphysema Pulmonary edema Löffler's syndrome / Eosinophilic pneumonia Respiratory hypersensitivity Allergic bronchopulmonary aspergillosis Hamman-Rich syndrome Idiopathic pulmonary fibrosis Sarcoidosis Vaping-associated pulmonary injury Obstructive / Restrictive Pneumonia / pneumonitis By pathogen Viral Bacterial Pneumococcal Klebsiella Atypical bacterial Mycoplasma Legionnaires' disease Chlamydiae Fungal Pneumocystis Parasitic noninfectious Chemical / Mendelson's syndrome Aspiration / Lipid By vector/route Community-acquired Healthcare-associated Hospital-acquired By distribution Broncho- Lobar IIP UIP DIP BOOP-COP NSIP RB Other Atelectasis circulatory Pulmonary hypertension Pulmonary embolism Lung abscess Pleural cavity / mediastinum Pleural disease Pleuritis/pleurisy Pneumothorax / Hemopneumothorax Pleural effusion Hemothorax Hydrothorax Chylothorax Empyema/pyothorax Malignant Fibrothorax Mediastinal disease Mediastinitis Mediastinal emphysema Other/general Respiratory failure Influenza Common cold SARS Coronavirus disease 2019 Idiopathic pulmonary haemosiderosis Pulmonary alveolar proteinosis
-
Vitamin K Deficiency
Wikipedia
External links [ edit ] Classification D ICD - 10 : E56.1 ICD - 9-CM : 269.0 MeSH : D014813 DiseasesDB : 13962 External resources eMedicine : med/2385 Patient UK : Vitamin K deficiency v t e Malnutrition Protein-energy malnutrition Kwashiorkor Marasmus Catabolysis Vitamin deficiency B vitamins B 1 Beriberi Wernicke–Korsakoff syndrome Wernicke's encephalopathy Korsakoff's syndrome B 2 Riboflavin deficiency B 3 Pellagra B 6 Pyridoxine deficiency B 7 Biotin deficiency B 9 Folate deficiency B 12 Vitamin B 12 deficiency Other A: Vitamin A deficiency Bitot's spots C: Scurvy D: Vitamin D deficiency Rickets Osteomalacia Harrison's groove E: Vitamin E deficiency K: Vitamin K deficiency Mineral deficiency Sodium Potassium Magnesium Calcium Iron Zinc Manganese Copper Iodine Chromium Molybdenum Selenium Keshan disease Growth Delayed milestone Failure to thrive Short stature Idiopathic General Anorexia Weight loss Cachexia Underweight v t e Conditions originating in the perinatal period / fetal disease Maternal factors complicating pregnancy, labour or delivery placenta Placenta praevia Placental insufficiency Twin-to-twin transfusion syndrome chorion / amnion Chorioamnionitis umbilical cord Umbilical cord prolapse Nuchal cord Single umbilical artery presentation Breech birth Asynclitism Shoulder presentation Growth Small for gestational age / Large for gestational age Preterm birth / Postterm pregnancy Intrauterine growth restriction Birth trauma scalp Cephalohematoma Chignon Caput succedaneum Subgaleal hemorrhage Brachial plexus injury Erb's palsy Klumpke paralysis Affected systems Respiratory Intrauterine hypoxia Infant respiratory distress syndrome Transient tachypnea of the newborn Meconium aspiration syndrome Pleural disease Pneumothorax Pneumomediastinum Wilson–Mikity syndrome Bronchopulmonary dysplasia Cardiovascular Pneumopericardium Persistent fetal circulation Bleeding and hematologic disease Vitamin K deficiency bleeding HDN ABO Anti-Kell Rh c Rh D Rh E Hydrops fetalis Hyperbilirubinemia Kernicterus Neonatal jaundice Velamentous cord insertion Intraventricular hemorrhage Germinal matrix hemorrhage Anemia of prematurity Gastrointestinal Ileus Necrotizing enterocolitis Meconium peritonitis Integument and thermoregulation Erythema toxicum Sclerema neonatorum Nervous system Perinatal asphyxia Periventricular leukomalacia Musculoskeletal Gray baby syndrome muscle tone Congenital hypertonia Congenital hypotonia Infections Vertically transmitted infection Neonatal infection rubella herpes simplex mycoplasma hominis ureaplasma urealyticum Omphalitis Neonatal sepsis Group B streptococcal infection Neonatal conjunctivitis Other Miscarriage Perinatal mortality Stillbirth Infant mortality Neonatal withdrawal
-
Congenital Afibrinogenemia
Wikipedia
External links [ edit ] Classification D ICD - 10 : D68.2 ICD - 9-CM : 286.3 OMIM : 202400 MeSH : D000347 DiseasesDB : 307 External resources MedlinePlus : 001313 eMedicine : ped/3042 v t e Disorders of bleeding and clotting Coagulation · coagulopathy · Bleeding diathesis Clotting By cause Clotting factors Antithrombin III deficiency Protein C deficiency Activated protein C resistance Protein S deficiency Factor V Leiden Prothrombin G20210A Platelets Sticky platelet syndrome Thrombocytosis Essential thrombocythemia DIC Purpura fulminans Antiphospholipid syndrome Clots Thrombophilia Thrombus Thrombosis Virchow's triad Trousseau sign of malignancy By site Deep vein thrombosis Bancroft's sign Homans sign Lisker's sign Louvel's sign Lowenberg's sign Peabody's sign Pratt's sign Rose's sign Pulmonary embolism Renal vein thrombosis Bleeding By cause Thrombocytopenia Thrombocytopenic purpura : ITP Evans syndrome TM TTP Upshaw–Schulman syndrome Heparin-induced thrombocytopenia May–Hegglin anomaly Platelet function adhesion Bernard–Soulier syndrome aggregation Glanzmann's thrombasthenia platelet storage pool deficiency Hermansky–Pudlak syndrome Gray platelet syndrome Clotting factor Hemophilia A/VIII B/IX C/XI von Willebrand disease Hypoprothrombinemia/II Factor VII deficiency Factor X deficiency Factor XII deficiency Factor XIII deficiency Dysfibrinogenemia Congenital afibrinogenemia Signs and symptoms Bleeding Bruise Hematoma Petechia Purpura Nonthrombocytopenic purpura By site head Epistaxis Hemoptysis Intracranial hemorrhage Hyphema Subconjunctival hemorrhage torso Hemothorax Hemopericardium Pulmonary hematoma abdomen Gastrointestinal bleeding Hemobilia Hemoperitoneum Hematocele Hematosalpinx joint Hemarthrosis
-
Isochromosome
Wikipedia
Consequently, there is partial trisomy of the genes present in the isochromosome and partial monosomy of the genes in the lost arm. [2] Contents 1 Nomenclature 2 Mechanism 2.1 Centromere misdivision 2.2 U-type strand exchange 3 Consequences 3.1 Turner syndrome 3.2 Neoplasia 4 References Nomenclature [ edit ] An isochromosome can be abbreviated as i(chromosome number and arm). ... Regardless of the chromosome involved in U-type exchange, the acentric fragment of the chromosome is lost, thus creating a partial monosomy of genes located in that portion of the acentric chromosome. [2] Consequences [ edit ] The most common isochromosome is the X sex chromosome . [4] Acrocentric autosomal chromosomes 13 , 14 , 15 , 21 , and 22 are also common candidates for isochromosome formation. [1] Chromosomes containing smaller arms are more likely to become isochromosomes because the loss of genetic material in those arms can be tolerated. [2] Turner syndrome [ edit ] Turner syndrome is a condition in females in which there is partial or complete loss of one X chromosome. This causes symptoms such as growth and sexual development problems. In 15% of Turner syndrome patients, the structural abnormality is isochromosome X, which is composed of two copies of the q arm (i(Xq)). [1] [2] A majority of i(Xq) are created by U-type strand exchange. ... Since the p-arm of the X chromosome contains genes that are necessary for normal sexual development, Turner's syndrome patients experience phenotypic effects. [3] Alternatively, the increase in dosage of genes on the q arm may be involved in a 10-fold increase in risk of i(Xq) Turner's patients developing autoimmune thyroiditis , a disease in which the body creates antibodies to target and destroy thyroid cells. [6] Neoplasia [ edit ] Neoplasia is uncontrolled cell growth, resulting in the creation of a tumour. ... Journal of Medical Genetics . 46 (10): 694–702. doi : 10.1136/jmg.2008.065052 . ^ Elsheikh, M.; Wass, J.A.H.; Conway, G.S. (2001). "Autoimmune thyroid syndrome in women with Turner's syndrome-the association with karyotype".TP53, IGHV1-12, MYC, MYCN, BCR, SETBP1, PML, RHOJ, KRAS, HTC2, SBDS, PEG10, SETMAR, RTL1, SLC45A2, SYK, TERT, TMX2-CTNND1, UROD, XIC, MOGS, TCL1A, SMC1A, RTL4, PPM1D, ENDOU, IMP3, AURKB, RECQL4, BCAR1, DLEU7, TCL1B, S100A10, SS18L1, RTL3, B3GAT1, IGHV1-17, ASXL1, EXOSC6, PAN3, AFP, RARA, RUNX1T1, DMD, CTNND1, CSF3, CSE1L, CD38, RUNX3, RUNX1, MAPK1, CAD, ATR, ATM, ARAF, AKT1, CRYBG1, EPHB2, ERBB2, FGF2, GH1, HLA-C, IGF1R, JAK2, KIT, LBR, LDHB, PAX1, PAX5, PIK3CA, PIK3CB, PIK3CD, PIK3CG, JAG1, TET18P
-
Congenital Hearing Loss
Wikipedia
Autosomal dominant congenital hearing loss can be attributed to such causes like Waardenburg Syndrome. Autosomal recessive hearing loss [ edit ] In autosomal recessive hearing loss, both parents who typically have normal hearing, carry a recessive gene . ... She can pass on the trait to males and female children, but usually only male children are affected. There are some genetic syndromes, in which hearing loss is one of the known characteristics. Some examples are Down syndrome (aneuploidy), Usher syndrome (autosomal recessive), Treacher Collins syndrome (autosomal dominant), Crouzon syndrome (autosomal dominant), and Alport syndrome (X-linked).GJB2, SLC26A4, KCNQ1, GJB6, KCNE1, POU3F4, MYO7A, TMPRSS3, USH2A, TMC1, CDH23, MYO15A, SMPX, KCNQ4, TECTA, LHFPL5, KCNQ3, SLC17A8, ATP2B2, PTGDS, CACNA1D, FGF3, STT3A, RCC1, LHFPL6, GIPC1, CSTB, CLDN14, COL9A3, MRPS7, CNC2, IL23A, ANKRD7, CLCNKB, ADGRV1, KCNE2, DBA2, SLC26A7, CDKN2D, LOXHD1, MARVELD2, CDKN2A, ILDR1, PTPRQ, CEACAM16, USH1C, ATG5, HINT1, EYA1, GATA2, KCNQ2, GAS1, KIF5A, KIT, MBL2, MYO6, NT5E, PAX3, F9, POU4F3, OTOF, PRKAR1A, CACNA1C, REG1A, REST, SLC8A1, ESRRG, USH1E, EDNRB, BSND, DFNB17, H3P13
-
Nephronophthisis 1
Omim
When nephronophthisis is combined with retinitis pigmentosa, the disorder is known as Senior-Loken syndrome (SLSN1; 266900); when it is combined with cerebellar vermis hypoplasia, the disorder is known as Joubert syndrome (JBTS1; 213300); and when it is combined with multiple developmental and neurologic abnormalities, the disorder is often known as Meckel-Gruber syndrome (MKS1; 249000). Because most NPHP gene products localize to the cilium or its associated structures, nephronophthisis and the related syndromes have been termed 'ciliopathies' (summary by Hoff et al., 2013). ... The microsatellite marker at the D2S160 locus gave a lod score of 4.78 at theta = 0.05 in 18 families with isolated NPH, whereas the same marker excluded linkage in the 4 families with associated ocular abnormalities (thus corroborating the view that Senior-Loken syndrome (266900) is a distinct entity). ... A similar association was not observed for 155 patients with Joubert syndrome (JBTS; see, e.g., 213300). Associations Pending Confirmation For discussion of a possible association between NPHP-related ciliopathies and variation in the IFT81 gene, see 605489.0001 and 605489.0002. ... INHERITANCE - Autosomal recessive GROWTH Other - Growth retardation GENITOURINARY Kidneys - Nephronophthisis - End stage renal disease - Tubular atrophy - Tubular basement membrane disintegration - Interstitial fibrosis - Corticomedullary renal cysts METABOLIC FEATURES - Polyuria - Polydipsia - Absence of hypertension HEMATOLOGY - Anemia LABORATORY ABNORMALITIES - Hyposthenuria (inability to concentrate urine normally) MISCELLANEOUS - Medial onset of end stage renal disease 13 years - Allelic to Senior-Loken syndrome 1 ( 266900 ) and Joubert syndrome 4 ( 609583 ) MOLECULAR BASIS - Caused by mutation in the nephrocystin 1 gene (NPHP1, 607100.0001 ) ▲ Close
-
Long-Chain 3-Hydroxyacyl-Coa Dehydrogenase Deficiency
Omim
Clinical Features Wanders et al. (1989) described sudden infant death syndrome (SIDS) in a 3-day-old infant caused by deficiency of long-chain 3-hydroxyacyl-CoA dehydrogenase. ... These maternal illnesses include the acute fatty liver pregnancy (AFLP) syndrome; hypertension or hemolysis, elevated liver enzymes, and low platelets (HELLP) syndrome; and hyperemesis gravidum. The AFLP syndrome is characterized by anorexia, nausea, vomiting, abdominal pain, and jaundice in the third trimester. Fulminant liver failure and death may occur. HELLP syndrome is more common and may represent the severe end of the spectrum of preeclampsia. In both syndromes, microvesicular fatty infiltration of maternal liver occurs, a pathologic picture similar to that in children with fatty acid oxidation defects.
-
Staphylococcal Infection
Wikipedia
Contents 1 Types 2 Coagulase-positive 3 Coagulase-negative 4 Causes 5 Signs and symptoms 6 Treatment 7 Etymology 8 Epidemiology 9 References 10 External links Types [ edit ] Main Staphylococcus aureus infections Type Examples Localized skin infections Stye and other small, superficial abscesses in sweat or sebaceous glands Subcutaneous abscesses ( boils ) around foreign bodies Large, deep infections ( carbuncles ) possibly causing bacteremia Folliculitis ; an infection of a hair follicle Ear infections [2] Diffuse skin infection Impetigo Cellulitis Deep, localized infections Acute and chronic osteomyelitis Septic arthritis Other infections Asthma and rhinitis [3] Acute [4] and chronic sinusitis Acute infective endocarditis Sepsis Necrotizing pneumonia Toxinoses Toxic shock syndrome Gastroenteritis Scalded skin syndrome Unless else specified in boxes, then reference is Lippincott's Illustrated Reviews: Microbiology (2007). [5] Other infections include: Closed-space infections of the fingertips, known as paronychia . ... S. aureus is also implicated [6] in toxic shock syndrome ; during the 1980s some tampons allowed the rapid growth of S. aureus, which released toxins that were absorbed into the bloodstream. Any S. aureus infection can cause the staphylococcal scalded skin syndrome , a cutaneous reaction to exotoxin absorbed into the bloodstream. ... Cellulitis commonly infects the lower legs, but can also, less commonly, affect the face and arms. Staphylococcus scalded skin syndrome – Staphylococcus scalded skin syndrome is caused by toxins produced when a staph infection gets too severe. ... External links [ edit ] Classification D MeSH : D013203 v t e Firmicutes (low- G+C ) Infectious diseases Bacterial diseases : G+ Bacilli Lactobacillales ( Cat- ) Streptococcus α optochin susceptible S. pneumoniae Pneumococcal infection optochin resistant Viridans streptococci : S. mitis S. mutans S. oralis S. sanguinis S. sobrinus S. anginosus group β A bacitracin susceptible: S. pyogenes Group A streptococcal infection Streptococcal pharyngitis Scarlet fever Erysipelas Rheumatic fever B bacitracin resistant, CAMP test +: S. agalactiae Group B streptococcal infection ungrouped Streptococcus iniae Cutaneous Streptococcus iniae infection γ D BEA +: Streptococcus bovis Enterococcus BEA +: Enterococcus faecalis Urinary tract infection Enterococcus faecium Bacillales ( Cat+ ) Staphylococcus Cg+ S. aureus Staphylococcal scalded skin syndrome Toxic shock syndrome MRSA Cg- novobiocin susceptible S. epidermidis novobiocin resistant S. saprophyticus Bacillus Bacillus anthracis Anthrax Bacillus cereus Food poisoning Listeria Listeria monocytogenes Listeriosis Clostridia Clostridium ( spore -forming) motile: Clostridium difficile Pseudomembranous colitis Clostridium botulinum Botulism Clostridium tetani Tetanus nonmotile: Clostridium perfringens Gas gangrene Clostridial necrotizing enteritis Finegoldia (non-spore forming) Finegoldia magna Mollicutes Mycoplasmataceae Ureaplasma urealyticum Ureaplasma infection Mycoplasma genitalium Mycoplasma pneumoniae Mycoplasma pneumonia Anaeroplasmatales Erysipelothrix rhusiopathiae Erysipeloid v t e Dermatitis and eczema Atopic dermatitis Besnier's prurigo Seborrheic dermatitis Pityriasis simplex capillitii Cradle cap Contact dermatitis ( allergic , irritant ) plants: Urushiol-induced contact dermatitis African blackwood dermatitis Tulip fingers other: Abietic acid dermatitis Diaper rash Airbag dermatitis Baboon syndrome Contact stomatitis Protein contact dermatitis Eczema Autoimmune estrogen dermatitis Autoimmune progesterone dermatitis Breast eczema Ear eczema Eyelid dermatitis Topical steroid addiction Hand eczema Chronic vesiculobullous hand eczema Hyperkeratotic hand dermatitis Autosensitization dermatitis / Id reaction Candidid Dermatophytid Molluscum dermatitis Circumostomy eczema Dyshidrosis Juvenile plantar dermatosis Nummular eczema Nutritional deficiency eczema Sulzberger–Garbe syndrome Xerotic eczema Pruritus / Itch / Prurigo Lichen simplex chronicus / Prurigo nodularis by location: Pruritus ani Pruritus scroti Pruritus vulvae Scalp pruritus Drug-induced pruritus Hydroxyethyl starch-induced pruritus Senile pruritus Aquagenic pruritus Aquadynia Adult blaschkitis due to liver disease Biliary pruritus Cholestatic pruritus Prion pruritus Prurigo pigmentosa Prurigo simplex Puncta pruritica Uremic pruritus Other substances taken internally: Bromoderma Fixed drug reaction Nummular dermatitis Pityriasis alba Papuloerythroderma of Ofuji